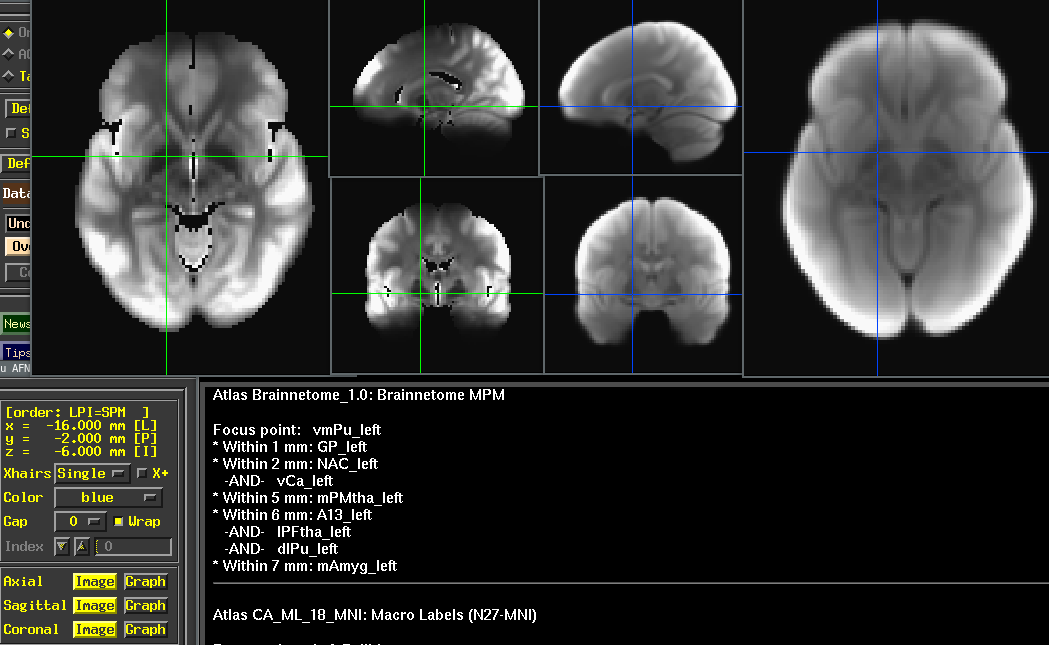
Time averaged T2*
Tip
For the 2026 manuscript, tat2 has been moved and renamed to dR2star.
Validation code for that is hosted in own repository tat2-validation
tat2 (code) wraps around 3dROIstats, 3dcalc, and 3dTstat to reduce 4D EPI BOLD data to a per-voxel (3D) measure (nT2*) that is inversely related to iron concentration.
This is used in
- Contributions of dopamine-related basal ganglia neurophysiology to the developmental effects of incentives on inhibitory control
- In vivo evidence of neurophysiological maturation of the human adolescent striatum
And is initially from Predicting Individuals' Learning Success from Patterns of Pre-Learning MRI Activity
The normalization is within the brain for each time point. Not all voxels in the volume, but only those that belong to the brain mask. Then we normalize so that the sum of all brain voxels is some fixed number, e.g., 10000. The number doesn't really matter.
Also see Relative Concentration of Brain Iron (rcFe).
Setup
See the setup instructions for all of lncdtools. Breifely, git clone https://github.com/lncd/lncdtools and add the new directory to your path.
The raw tat2 script can stand alone, but will uses other lncdtools scripts if avaiable -- namely gitver. It's also a lot easier to fetch and track updates when the script is within source control. You get that when you clone the repo.
Usage
see tat2 --help
Simple
Will combine all of the matching *preroc_bold.nii.gz (presumably multiple 4D time series fMRI data) into a single 3D image derive/tat2.nii.gz
fmriprep
Very limited fmriprep support is provided. More support is offered by dR2star.
export FD_THRES=0.3
tat2 -fmriprep /path/to/fmriprep-deriv/
# will make a tat2star file for each func folder found.
# e.g. /path/to/fmriprep-deriv/sub-1/ses-1/func/sub-1_ses-1_desc-preproc_tat2star.nii.gz
With options and relative paths
See https://regex101.com/r/bM6p7X/1 for a visual representation and playground of the regular expressions explored below.
The reference region mask and/or censor files may be in another folder or named specific to a participant's visit and run input file.
-mask_rel and -censor_rel support s/search/replace/ regular expressions to transform the input *.nii.gz name into a matching motion or censor file.
Imagine a somewhat pathological file organization like
├── censor_files
│ └── sub-1_ses-2_run-1
│ └── preproc_censor-fd0.3.1D
│ └── sub-1_ses-2_run-2
│ └── preproc_censor-fd0.3.1D
└── func
└── sub-1_ses-2_run-1_desc-preproc_bold.nii.gz
└── sub-1_ses-2_run-1_desc-brain_mask.nii.gz
...
└── sub-1_ses-2_run-2_desc-preproc_bold.nii.gz
└── sub-1_ses-2_run-2_desc-brain_mask.nii.gz
tat2 can accommodate matching pairing each run to the appropriate file using regular expression search and replace.
sub_ses="sub-01/ses-01"
censor_regex='s/.*func\/(.*)_desc-preproc_bold.nii.gz/censor_files\/\1\/preproc_censor-fd0.3.1D/'
tat2 \
-output "deriv/$sub_ses/func/${sub_ses//\//_}_space-MNI152NLin2009cAsym_tat2.nii.gz" \
-mask_rel 's/desc-preproc_bold.nii.gz/brain_mask.nii.gz/' \
-censor_rel "$censor_regex" \
-median_time \
-median_vol \
-no_voxscale \
-verbose \
deriv/$sub_ses/func/*space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
- sibling mask files are matched using
-mask_rel 's/desc-preproc_bold.nii.gz/brain_mask.nii.gz/'func/sub-1_ses-2_run-1_desc-preproc_bold.nii.gz # becomes func/sub-1_ses-2_run-1_desc-brain_mask.nii.gz- searches each input file for
desc-preproc_bold.nii.gz - and replaces that with
brain_maks.nii.gz
- searches each input file for
- to match censor files across directories,
censor_regexusess/.*func\/(.*)_desc-preproc_bold.nii.gz/censor_files\/\1\/preproc_censor-fd0.3.1D/). Also see https://regex101.com/r/bM6p7X/1func/sub-1_ses-2_run-1_desc-preproc_bold.nii.gz # becomes censor_files/sub-1_ses-2_run-1/preproc_censor-fd0.3.1D- searches each input file for
.*func\/(.*)_desc-preproc_bold.nii.gz, where\/"escapes" the directory slash, escape to distinguish it from the search-replace deliminator ins///
- where
(.*)captures the matching part to reuse as\\1in - the replacement like
censor_files\/\1\/preproc_censor-fd0.3.1D
- searches each input file for
Also see issue#5.
Preprocessing
We slice-time and motion correction, skull strip, despiking (wavelet), and warp to MNI before running tat2.
The validation manuscript (tat2-validation) also uses ABCD minimally preprocessed inputs.
Caution
Notably, smoothing is not included in datasets input to tat2.
Pipeline

Example
Example mean tat2 images 1
Comparisons
Permutation of tat2 calls were compared against R2 acquisitions.
-vol_median is likely the appropriate normalization.
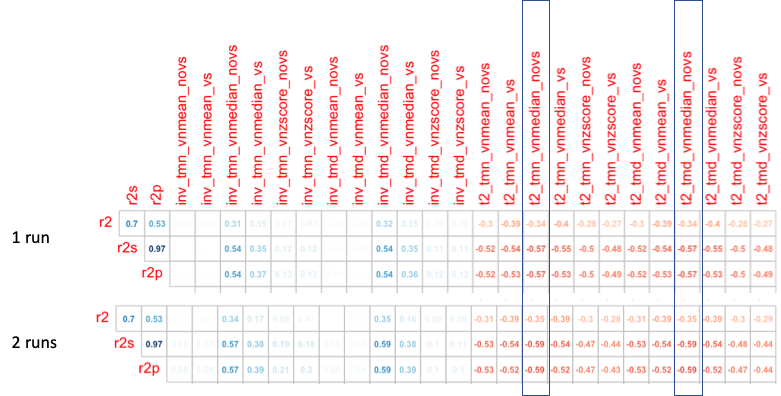
Correlation with R2
tat2 is negatively correlated R2* 2